DNA microarrays are highly effective detection systems for nucleic acids. They can be used to detect and discriminate against infectious agents. This is particularly important for the diagnostics of diseases where a highly parallel detection and discrimination of pathogens is required.
One focus of the institute is the rapid identification of pathogens and the detection of antibiotic resistance using DNA arrays. For more than 10 years we have been developing DNA microarrays for customers from companies, clinics and research institutions, individually adapted to specific indications. The infrastructure in the institute enables the complete production, from the establishment of multiplex PCR and amplification of pathogen targets, probe design and immobilization of probes by contact printing to the hybridization of target DNAs for fluorescence-based identification of pathogens.
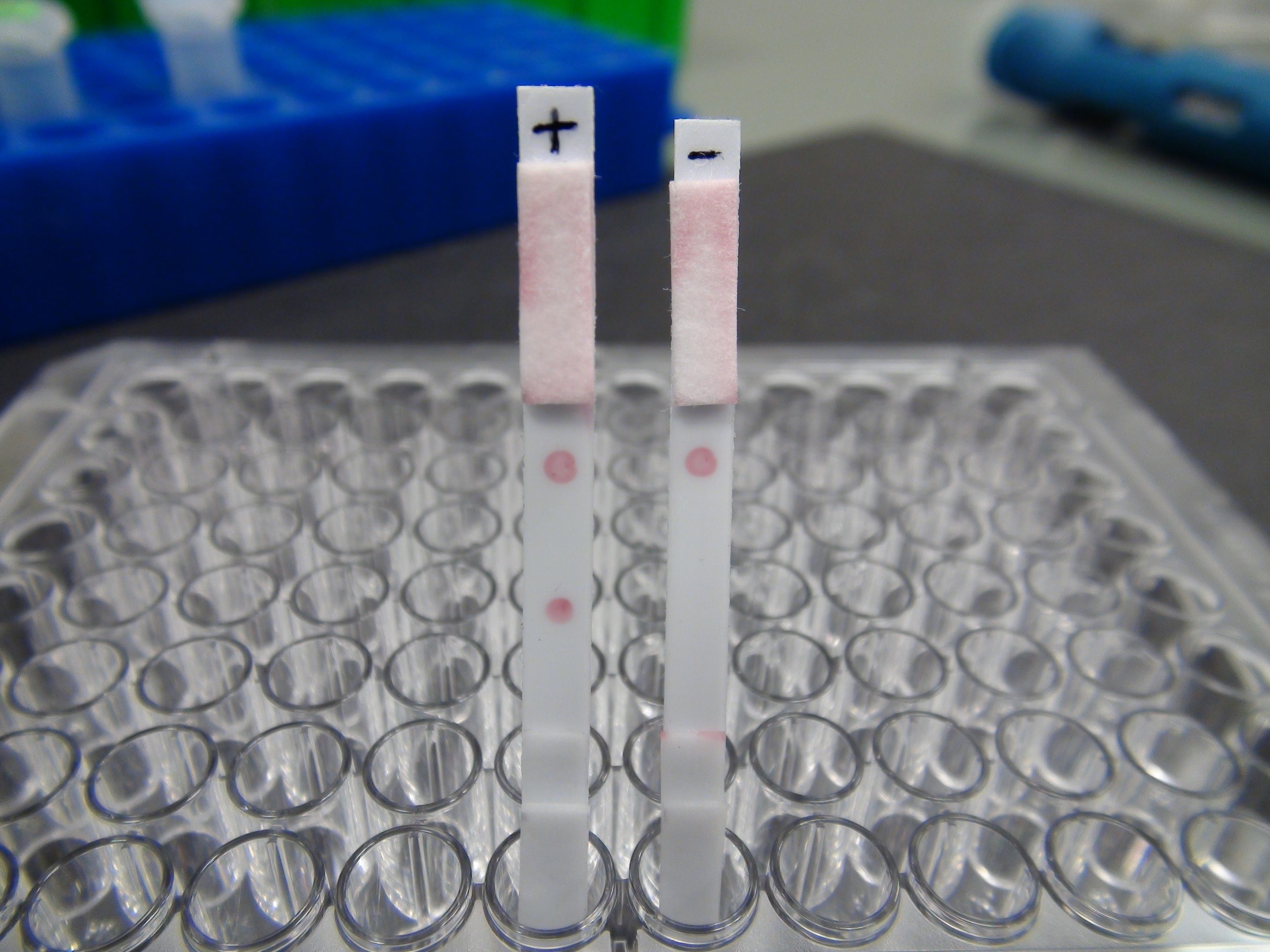
The basic principle of a microarray is the ordered, point-like immobilization of up to thousands of short defined DNA probes on a solid support. Using hybridization, the immobilized DNA probes which are complementary to the pathogen target sequence allow to capture the genomic target of respective pathogens. To this end, target sequences of pathogens present in a patient sample are amplified by PCR and detected by binding to specific immobilized probes. Simultaneously present probes, each specific for a particular pathogen, allow the parallel analysis of pathogens or resistances. In this way, complex causes of disease can be detected and discriminated against in just a few hours in a single sample and and a single diagnostic approach.
For example, in the BMBF-funded project Fungal Yeast Identification, we have developed a fully integrated lab-on-a-chip system for the rapid determination of approximately 50 yeast and fungi infections in cooperation with the companies Euroimmun, Multi Channel Systems MCS and Bosch as well as the Heart and Diabetes Centre North Rhine-Westphalia. In addition, a DNA microarray platform was developed in close cooperation with GATC Biotech AG, with which sepsis pathogens (bacteria and fungi) as well as their relevant resistances to antibiotics can be detected and identified in parallel. On behalf of Immundiagnostik AG Bensheim, we have developed DNA-based microarrays for the highly parallel diagnosis of fungal infections and sexually transmitted infectious diseases caused by fungi, bacteria, viruses or protozoa.
 Fraunhofer Institute for Interfacial Engineering and Biotechnology IGB
Fraunhofer Institute for Interfacial Engineering and Biotechnology IGB